FuncNet
FuncNet is a distributed protein function comparison pipeline, funded by the European Union's EMBRACE Network of Excellence, and developed in partnership with the ENFIN project.
This page is no longer maintained. See funcnet.eu instead.
Aims
The objective of FuncNet is to provide an open platform for the computational prediction and analysis of protein function.
It is designed to answer questions like:
Given one set of proteins which are known to share a particular biological function…
… which of these other proteins also share that function?
A good example of this is the prediction of proteins involved in the formation and activity of the mitotic spindle. Since a set of known spindle proteins already exists (Sauer et al. 2005), FuncNet can be used to predict whether uncharacterized or partially-characterized proteins also belong in this set, by aggregating pairwise functional similarity predictions between query and reference proteins.
Implementation
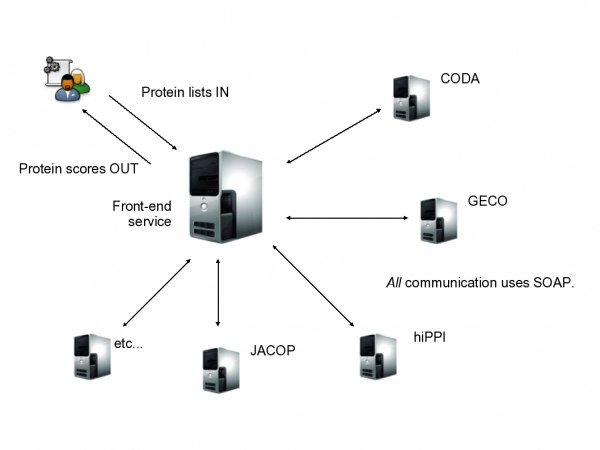
FuncNet is an open architecture on which multiple prediction algorithms can be queried in parallel in order to provide higher-quality results. Each predictor is made available via a SOAP web service using a standardized WSDL interface. This means that every FuncNet prediction service is functionally interchangeable – they can all be invoked via the same message format (described below), and you only need to change the endpoint URL and the service and port names. The current template WSDL for FuncNet services is here.
However, a better way to submit queries is via a front-end service which is responsible for forwarding the request to each of the predictors in parallel. On receipt of the results, it uses Fisher's unweighted method to integrate the various predictors' responses into a single prediction for each query protein:
Of course, users can submit queries directly to the individual predictors, although the strength of FuncNet comes from its ability to combine the predictions of multiple algorithms which use distinct methods and sources of evidence.
Predictors
There are currently five prediction algorithms online, using various different sources of evidence:
- CODA (hosted at UCL): evolutionary relatedness based on domains found together in other species
- engineDB (hosted at CNR-ITB): detection of functionally analogous proteins via GO annotations
- GECO (hosted at UCL): correlated patterns of gene expression from microarray experiments
- hiPPI (hosted at UCL): homology-based inheritance of protein-protein interactions from public databases
- JACOP (hosted at SIB): unsupervised clustering and classification based on detection of homologous sub-sequences
Usage
FuncNet queries can be submitted from any SOAP client which supports the 'document/literal wrapped' style. Almost all modern SOAP toolkits support this model. The interface is intentionally very simple so databinding shouldn't be a problem, regardless of what programming language you use, and of course you can 'roll your own' XML if you prefer.
To get you started, we've provided some example libraries and scripts (see Links below) and will be adding more samples in different languages over time. Feel free to send us your own!
If you want to try out FuncNet without doing any coding, download soapUI. This is a very handy Java tool for testing web services. Choose 'New WSDL Project' from the File menu, paste in the URL of a FuncNet WSDL (see below), and it'll generate the appropriate request templates for you. Then you can insert some UniProt primary accessions in the <p> fields (see below), submit the query by pressing the green 'play' button, and wait for some results.
We are working on integrating FuncNet into the EnCORE web services framework too – check back here for further news.
NB FuncNet only understands UniProt primary accessions, and is currently limited to human proteins.
Predictor request format
The standard format for a request to a FuncNet prediction service (without the SOAP wrappings) looks like this:
<funcnet:ScorePairwiseRelations xmlns:funcnet="http://cathdb.info/FuncNet_0_1/">
<proteins1>
<!-- 1 or more UniProt primary accessions: -->
<p>Q8NFN7</p>
</proteins1>
<proteins2>
<!-- 1 or more UniProt primary accessions: -->
<p>Q8NF37</p>
</proteins2>
</funcnet:ScorePairwiseRelations>
By convention, proteins1 is the list of query proteins (unknown function) and proteins2 is the list of reference proteins (known function).
NB Requests to JACOP must provide at least three reference proteins.
Predictor response format
<funcnet:ScorePairwiseRelationsResponse xmlns:funcnet="http://cathdb.info/FuncNet_0_1/">
<!-- One or more score tuples: -->
<s>
<p1>Q8NFN7</p1>
<p2>Q8NF37</p2>
<rs>65.20529</rs>
<pv>.140766</pv>
</s>
</funcnet:ScorePairwiseRelationsResponse>
(The numbers in this example are made up and don't come from any real prediction service.)
p1 = an accession from the proteins1 list
p2 = an accession from the proteins2 list
rs = raw score from the prediction algorithm (not comparable between algorithms)
pv = p-value for the prediction
The p-value is formally defined as the probability that a random pair of proteins from the human genome would score equal to or higher than this pair using the same prediction algorithm. You can consider this as a test of significance at whatever cutoff you see fit (⇐0.05 is usually a safe bet).
The maximum number of scores that can be returned by a predictor = |proteins1| * |proteins2| and the minimum is zero. This is due to data sparsity and other factors; some of the predictors don't know anything at all about the relationship between a given pair of proteins and therefore won't even give them a low score. For example, GECO uses correlated patterns of expression in microarray experiments, and some genes just aren't commonly used on arrays, meaning it can't draw any conclusions about their products.
NB For performance purposes, many of the prediction services don't check that the accession codes supplied are genuine and from humans. This is the responsibility of the user. Unknown accessions will just be quietly ignored.
Partners
The current release of FuncNet is a collaboration between the Gene3D-BioMiner and CATH teams at University College London, Heinz Stockinger and Marco Pagni at the Swiss Institute of Bioinformatics, Andreas Gisel at ITB-CNR in Bari, and Juan Ranea and Ian Morilla at the University of Malaga.
It is co-ordinated by Andrew Clegg under the supervision of Christine Orengo.
There are two other EMBRACE groups involved in project whose contributions are in progress, the Valencia group at CNIO in Madrid, and the Brunak group at DTU-CBS in Lyngby. In addition, we have recently been joined by the Barton group in Dundee who are members of the ENFIN project.
Our EMBRACE liaison is Erik Bongcam-Rudloff at Uppsala University. We are grateful for the technical assistance of ENFIN's Florian Reisinger.
Current Status
The five prediction algorithms listed above are up and running individually (see Links). Feel free to submit queries to them.
The front-end service (including the statistical integration of results) is still under testing, and not yet available to the public.
NB In the CODA, GECO and hiPPI services, the Raw Score (rs) field for each prediction currently returns zero for every result. This is because we haven't yet imported these values into our database. However, the p-value is actually a more informative measure, since raw scores are not comparable between predictors. The front-end service only considers p-values when integrating scores from multiple predictors.
Links
WSDL files for web services
- CODA using CATH domains –
CodaCathService - CODA using Pfam domains –
CodaPfamService - engineDB –
engineDBService - GECO –
GecoService - hiPPI –
HippiService - JACOP –
JacopService
Because of the way the CXF web service toolkit generates production WSDLs from source, the CODA, GECO and hiPPI WSDLs are actually the same (each one contains the details for all of them).
The standard template from which they all derive is here.
The draft WSDL for the front-end service is also available, although the service doesn't work yet.
Client tools
- Perl module on CPAN
To install this library, type
perl -MCPAN -e 'install WebService::Cath::FuncNet'
If you need help with this process, particularly if you don't have root on your machine and some of the new dependencies default to needing root access, have a look at this tutorial.
This script shows very simply how to access one of the predictors. The full CPAN library is much more capable and well-documented, but this script shows how little effort it actually is.
You will need to install the XML::Compile module yourself if you download this (the CPAN library handles dependencies for you). Also you'll need to download the WSDL for the service you want to query (the CPAN library also does this for you).
Publications and documentation
- Gene analogue finder: a GRID solution for finding functionally analogous gene products (Tulipano et al., 2007)
- Prediction of new mitotic spindle proteins (Orengo, 2008, PDF presentation)
- Gene3D, Orthology and Homology-Based Inheritance of Protein-Protein Interactions (Yeats, 2008, PPT presentation)
Homepage
- Quick link to this page: http://funcnet.eu
Sponsors and related sites
Contact
Email Andrew Clegg with any enquiries, feedback or problem reports: spamproof@funcnet.eu but replace 'spamproof' with 'info'.