How To Find a Particular Domain
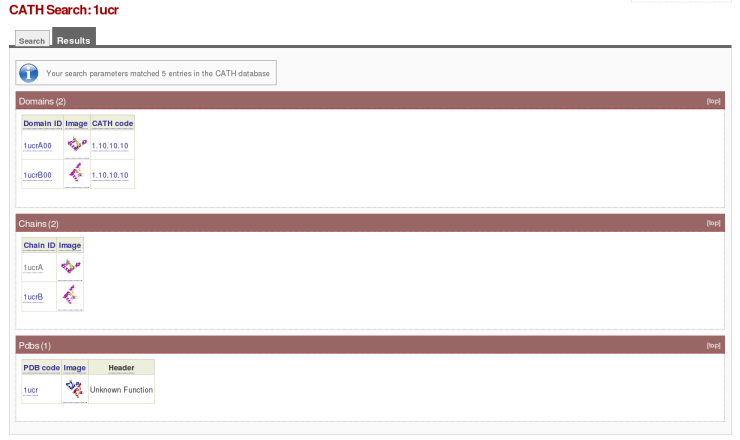
To find a particular domain, use either the CATH domain ID, CATH chain ID or the PDB code as the search term. The search term can be entered into the 'Quick Search' box in the top right-hand corner of any CATH page, or into the 'Search by ID/Keywords' box on the 'Search CATH by ID/sequence/text' page accessible from the CATH homepage. Using the protein '1ucr' as an example, a search should come up with the following:
Here you can find the following information:
- The protein consists of two domains, '1ucrA00' and '1ucrB00.' Both of these domains belong to the same homology family, '1.10.10.10.' To discover other domains that belong to this homology family, simply click on either one of the two given CATH codes
- '1ucr' is a dimer. Each of the chains within the dimer has its own unique chain ID (i.e. '1ucrA' and '1ucrB')
- The PDB code for the protein and the protein's function (which in this case is unknown)
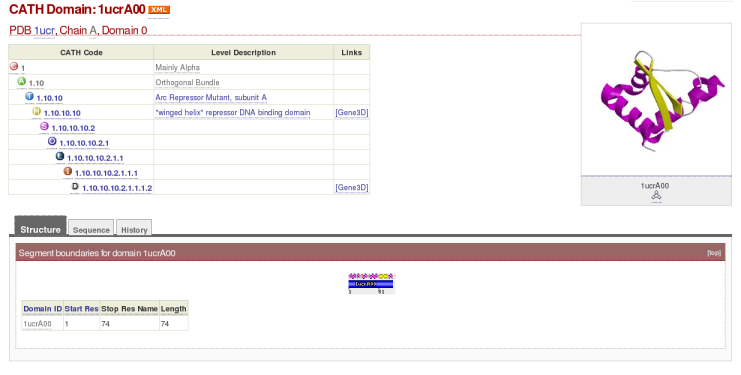
Clicking on a domain ID, e.g. '1ucrA00,' will redirect you to an information page about that domain. The information on this page has been broken down into three sections: 'Structure,' 'Sequence' and 'History.' The first section of the '1ucrA00' information page, the 'Structure' section, looks like this:
Here you can find the following information:
- The domain's unique CATH domain ID, as well as the CATH code, and the sequence family that the domain has been assigned to. Information concerning other domains assigned to the same level of CATH code can be obtained simply by clicking on the desirable level of the CATH code
Beneath the 'Structure' tab:
- The start and stop residue of the domain as well as the length of the domain can be found in the table 'Segment Boundaries'
- The 'Chopping Figure' given in this tab gives an indication of the domain's location within the rest of the chain. The chain given in this example is a whole chain domain (WCD): the entire chopping figure is a single colour.
An example of a chopping figure of a multi-domain chain is '1e4fT,' which looks like this:
Each colour represents a separate domain. Above the rectangular box representation of the domains are yellow and pink discontinuous irregular shapes. These shapes represent the secondary structure, alpha-helices (pink) and beta-strands (yellow), and their positions within the chain. The '1ucrA00' chopping figure shows that this domain consists of 5 alpha-helices and 3 beta-strands.
Beneath the 'Sequence' tab:
- The 'ATOM sequence,' which has been generated directly from the ATOM entry in the PDB file, and reflects the residues that are observed in the PDB structure
- The 'COMBS Sequence,' which tries to provide the entire residue sequence for that structure
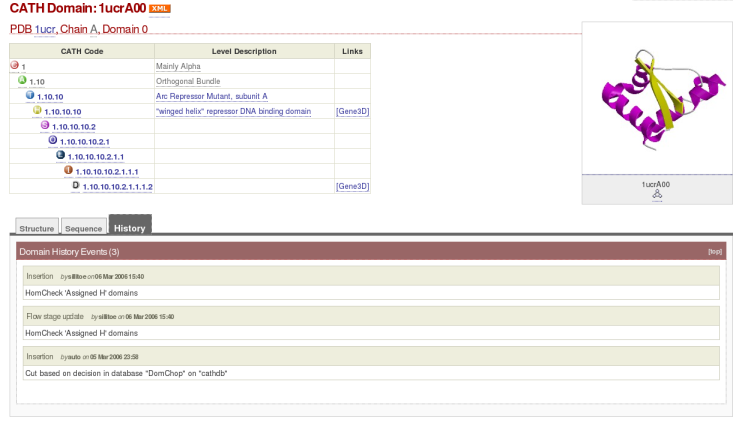
The final tab, 'History,' details the passage of the domain through CATH, including records concerning the chain chopping and assignment of the domain to its homologous superfamily. For 1ucrA00, the 'History' entry looks like this:
Links to Other Tutorial Pages
| Page | Description |
|---|---|
| browse_the_cath_hierarchy | How to Browse the Four Major Clusters in the CATH Hierarchy It is possible to browse the four major levels of the CATH hierarchy by clicking on 'Browse CATH from the top of the hierarchy,' which is located on the CATH start page under the heading 'Links for Researchers.' This will take you to a page that looks like this: |
| text_search | How To Perform a Text Search It is possible to search the CATH database using any search word. For example, the word may be of functional origin, such as 'chaperone,' or structural, such as 'helix'. The search word can be entered in the 'Quick Search' box in the top right-hand corner of the screen, or into the 'Search by ID/Keywords' box on the 'Search CATH by ID/sequence/text' page accessible from the CATH homepage. The search word 'lysine' will generate a result page that looks like this: |